 Evolution
Evolution
Stylus‘s Evolutionary Simulation Aims to Bridge Gap Between Real World and Artificial
In 2008, Biologic Institute director Doug Axe co-published an article in PloS One describing Stylus, a computer program created to simulate Darwinian evolution. Many computer simulations that purport to simulate Darwinian evolution have deficiencies: Pro-ID scientists like William Dembski or Robert Marks have shown how programs smuggle in information such that they are pre-directed to evolve their targets. Axe’s work demonstrates that such programs typically evolve solutions to artificial, rather than real-world problems. A new peer-reviewed paper in BIO-Complexity by Axe, Philip Lu, and Stephanie Flatau explains that the “functions” of the digital organisms in these simulations are often divorced from real-world meaning. They designed Stylus to present a more accurate picture:
The motivation for Stylus was the recognition that prior models used to study evolutionary innovation did not adequately represent the complex causal connection between genotypes and phenotypes.
(Douglas D. Axe, Philip Lu, and Stephanie Flatau, “A Stylus-Generated Artificial Genome with Analogy to Minimal Bacterial Genomes,” BIO-Complexity, Vol. 2011(3).)
Stylus aims to corrects these deficiencies by simulating Darwinian evolution in a manner that more accurately reflects the biological relationship between genotype and phenotype. It is also more realistic because it solves real-world problems. As the paper explains, “Functional specificity therefore has a structural basis in the Stylus world, just as it does in the real world.” Stylus manipulates digital objects that have real-world meaning: the targets of evolution in Stylus are Chinese characters. As the paper explains:
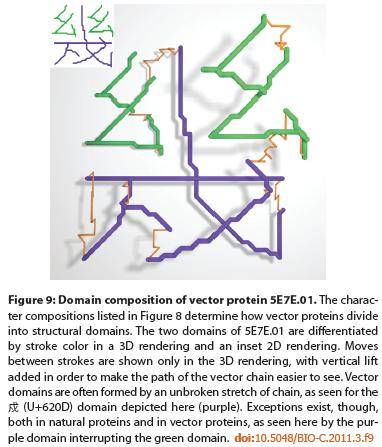
These translation products, called vector proteins, are functionless unless they form legible Chinese characters, in which case they serve the real function of writing. This coupling of artificial genetic causation to the real world of language makes evolutionary experimentation possible in a context where innovation can have a richness of variety and a depth of causal complexity that at least hints at what is needed to explain the complexity of bacterial proteomes.
These characters not only have real-world meaning, but their function-related shapes bear interesting analogies to proteins. An additional similarity between Chinese characters and proteins is that just as protein domains are re-used throughout many proteins, so particular shapes, called “strokes,” are found commonly throughout Chinese characters.
Sequence, Structure, and Function
Basic to life is an information conversion, where the information carried in genes (the genotype) is converted into an organism’s observable traits (the phenotype). Those biological structures then perform various functions. Another way of framing this information conversion is therefore:
sequence → structure → function
Axe, Lu and Flatau explain that many previous computer programs attempting to simulate evolution achieve part of this conversion, but not the whole thing.
For example, Conway’s famous Game of Life starts with a structure, and in some instances that structure can perform a function. But there is no sequence involved in the conversion. Avida starts with a sequence of programming commands, and when successful performs certain logic functions. But in Avida there is no structure to mediate between sequence and function.
Stylus, on the other hand, is more advanced in that it simulates the full sequence → structure → function information transfer. It does this by starting with a programmed genome. As the paper explains:
[The] Stylus genome encodes a special kind of text, namely, one that describes how to decode the genome. That is, the desired genome will encode a sequence of Chinese characters (in the form of vector proteins) that tells a reader of Chinese how Stylus genes are translated into vector sequences, and how those sequences are processed to make readable vector proteins.
The paper explains: “What Stylus offers that no other model offers, to our knowledge, is an artificial version of gene-to-protein genetic causation that parallels the real thing.”
Simulating Translation
Of course, if you’re going to simulate Darwinian evolution, then it would make sense to try to program something like the translation process. Though no analogy is perfect, Stylus makes a strong attempt to accomplish this.
At the sequence level, the Chinese characters are encoded by strings of four familiar letters: A, C, T, and G. And just like in biology, these letters form triplets, or codons that map to vectors, which combine to form the structure of the characters. Much like codons in the genetic code in biology encode amino acids, the codons of Stylus code for directional vectors that are used to draw the Chinese characters. The “genetic code” of Stylus is portrayed in the diagram below from the paper:
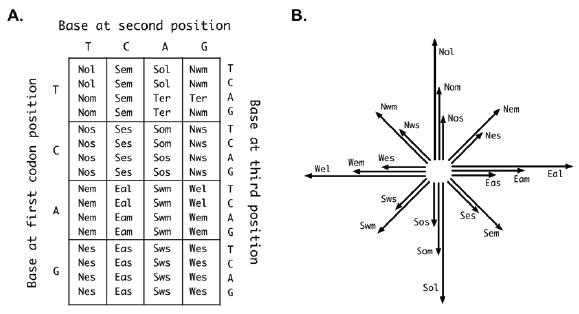
In Part A, various three-letter combinations of A, C, T, and G, encode various codons which are used to signify the “vector proteins,” which ultimately combine to form Chinese characters. As you can seen in the diagram, just as there are 20 amino acids that join to form proteins, Stylus uses 20 vectors to draw Chinese characters. The way these vectors can combine to draw a Chinese chracter is seen in the diagram from the paper below:
There are other important ways that Stylus‘s simulation parallels biological reality:
With these single-character representations of the twenty vectors established, the next step is to devise a compact way of specifying the codons that encode them. For this, we chose to elaborate on the concept of genes in the Stylus world. … Stylus genes depend on upstream sequence in order for translation to occur, as do real bacterial genes. Analogous to the Shine-Dalgarno sequence found on bacterial mRNAs, Stylus genes require an AGG sequence separated from the ATG start codon by twelve bases, as depicted. (internal citations omitted)
Evolving Chinese Characters
In the world of Stylus, a Chinese character is like a protein. So how can we determine if a functional “protein” has evolved? According to the paper, “At the core of Stylus software is an algorithm that quantifies the likeness of a given vector protein to a specified Chinese character.” This complicated algorithm is described as follows:
Stylus endows these graphical constructs with interesting similarities to their molecular counterparts by uncovering and exploiting a pre-existing analogy — the analogy between the set of characters used in Chinese writing and the set of protein structures used in life. Specifically, vector proteins are drawn objects that may function as legible Chinese characters if they are suitably formed. … Stylus is unique in its use of real function that maps well to molecular biology. It therefore represents a significant advance in the field of evolutionary modeling. (internal citations omitted)
Obviously, in a living bacterium, there are many hundreds if not thousands of genes. Since there are so many characters in the Chinese alphabet, again this serves as a useful way to model biology. The Stylus “proteome” thus consists of 223 Chinese characters which, much like protein families, group into 87 homologous groups.
But can these Chinese character groups, which have many qualities that parallel real-world protein families, evolve by random mutation and natural selection? That’s the sort of question the creators of Stylus hope to answer.
Great, but What Can We Do with It?
The paper presents a set of Chinese characters that can be used for simulating the evolutionary process in the Stylus world. The results of such simulations will probably be fleshed out in future papers. But the current paper leaves us with a strong sense of where this is all heading:
Evolutionary causation is intrinsically tied to the relationship between genotype and phenotype, which depends on low-level genetic causation. It follows that evolutionary explanations of the origin of functional protein systems must subordinate themselves to our understanding of how those systems operate. In other words, the study of evolutionary causation cannot enjoy the disciplinary autonomy that studies of genetic causation can.
In view of this, the contribution of Stylus is to make evolutionary experimentation possible in a model world where low-level genetic causation has the essential role that it has in the real world. Combined with the free Stylus software, the complete Stylus genome made freely available with this paper paves the way for analogy-based studies on a wide variety of important subjects, many of which are difficult to study by direct experimentation. Among these are the evolution of new protein folds by combining existing parts, the optimality and evolutionary optimization of the genetic code, the significance of selective thresholds for the origin and optimization of protein functions, and the reliability of methods used for homology detection and phylogenetic-tree construction.
There probably will never be a perfect computer simulation of biological evolution, but Stylus brings new and improved methods to the field of evolutionary modeling. This tool will help those interested in testing the viability of Darwinian claims to assess whether complex features can be created by random mutations at the molecular level.

