 Evolution
Evolution
 Intelligent Design
Intelligent Design
Dolphins and Porpoises and…Bats? Oh My! Evolution’s Convergence Problem
I have recently been reading George McGhee’s Convergent Evolution: Limited Forms Most Beautiful. McGhee’s book is a gripping read, and it favorably cites the work of both Michael Denton and Douglas Axe, ID-friendly scientists well known to readers of ENV. The book documents a multitude of cases of convergent evolution (homoplasy), the phenomenon of repeated evolution. When similarity is thought to have arisen by means of common ancestry, the features in question are said to be “homologous.” When similarity is thought to have arisen by means other than common ancestry, the features are said to be “analogous.”
 Those who subscribe to universal common ancestry interpret biological similarity of sequence, structure and anatomy as resulting from descent with modification from a common ancestral source. ID proponents who question common ancestry typically interpret biological similarity as resulting from a common blueprint. Is there a way to evaluate which of these two competing hypotheses better fits the evidence?
Those who subscribe to universal common ancestry interpret biological similarity of sequence, structure and anatomy as resulting from descent with modification from a common ancestral source. ID proponents who question common ancestry typically interpret biological similarity as resulting from a common blueprint. Is there a way to evaluate which of these two competing hypotheses better fits the evidence?
If you take such similarity as pointing to common descent, then you would expect to see it exhibiting a nested hierarchical distribution, the more seamless the better. In other words, the patterns of distribution of this similarity ought to mutually corroborate a single family tree. Sure, there might be occasional deviations from that tree, the results of phenomena such as incomplete lineage sorting. One would not expect to see the pervasive occurrence of a high degree of similarity — what would normally be regarded as “homology” — that decidedly cannot be accounted for within the framework of common descent. Yet that is in fact what we do observe.
Perhaps the most impressive examples of convergent evolution are those that occur at the molecular level — that of amino acid or even nucleotide sequences. Assessing the pervasiveness of convergence at the nucleotide sequence level is difficult, however, owing to a current lack of available data. Indeed, Castoe et al. (2010) point out that “complete vertebrate nuclear genomes still number in the tens, not hundreds.”
Even so, the little data that we do have yield startling results. One example of such convergence, documented by Zhang (2006), is the case of the pancreatic RNase in the African colobine monkeys and Asian colobine monkeys (thought to have diverged some 13 million years ago). Three identical nucleotide substitutions occurred in the DNA coding for these enzymes in the two lineages. This resulted in a lowering of the pH of the enzyme’s maximum ribonucleolytic activity from 7.4 to 6.3, allowing the enzyme to work in more acidic conditions than normal, making it perfectly suited for the small intestine.
Cuevas et al. (2002) have furthermore documented, in retroviruses, the occurrence of molecular convergences in 12 variable sites in independent lineages. Some of these convergent mutations even took place in intergenic regions (changes in which are normally thought to be selectively neutral) and also in synonymous sites. The authors note that this is fairly widely observed among HIV-1 virus clones in humans and in SHIV strains isolated from macaques, monkeys and humans.
Mitochondrial DNA data has revealed even more astonishing results. Castoe et al. (2009, 2010), for example, report the occurrence of 44 parallel amino acid substitutions in all 13 mitochondrially encoded oxidative phosphorylation metabolic proteins in the distantly related snakes and agamid lizards. Their 2009 paper notes:
These results indicate that nonneutral convergent molecular evolution in mitochondria can occur at a scale and intensity far beyond what has been documented previously, and they highlight the vulnerability of standard phylogenetic methods to the presence of nonneutral convergent sequence evolution.
Stern and Orgogozo (2009) examine 350 different mutations responsible for phenotypic variation and conclude that “more than half of these represent cases of parallel genetic evolution.” Moreover,
Gene function explains part but not all of the observed pattern of parallel genetic evolution. In several cases, parallelism has been observed even though mutations in a large number of genes can produce similar phenotypic changes. For example, although more than 80 genes regulate flowering time, changes in only a subset of these genes have produced evolutionary changes in flowering time. Hundreds of genes regulate the pattern of fine epidermal projections, called trichomes, on Drosophila melanogaster larvae. But only one gene, called shavenbaby, has evolved to alter larval trichome patterns between Drosophila species, and this gene has accumulated multiple evolutionarily relevant mutations. [internal citations omitted]
It is suggested that “hotspot genes” such as shavenbaby exist because,
In the entire regulatory network governing development of the Drosophila embryo, only shavenbaby, with its specialized function to rally the entire module of trichome morphogenesis, can accumulate mutations that alter trichome patterns without disrupting other developmental processes.
Protein amino acid sequences also exhibit pervasive convergence. One of my favorite examples is the convergent evolution of the motor protein Prestin in echolocating bats and cetaceans. Take a look at the following figure, excerpted from Liu et al. (2010).
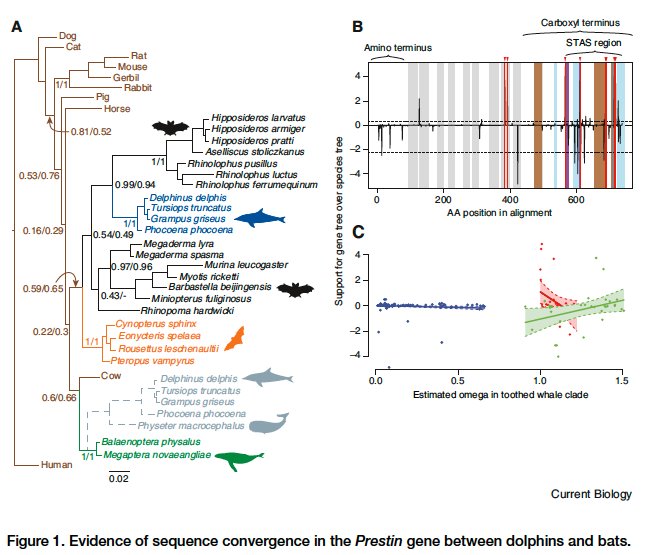
The paper reports that echolocating dolphins “group with echolocating bats in a phylogenetic tree of Prestin.” Jones (2010) states that,
…dolphins and porpoises share at least 14 derived amino acid sites in prestin with echolocating bats, including 10 shared with the highly specialized CF bats. Consequently, dolphins and porpoises form a sister group to CF bats in a phylogenetic analysis of prestin sequences (Figure 1). This finding is arguably one of the best examples of convergent molecular evolution discovered to date, and is exceptional because it is likely to be adaptive, driven by positive selection.
Another example is reported by Robson et al. (2000), who document that 28 of the amino acids in the oothecin protein of the cockroach are in exactly the same order as those in the lampry lamprin protein. The authors state that “sequence similarities between lamprin and oothecin, which share a 28/30 amino acid sequence identity, may represent one of the best examples of primary sequence convergence so far identified.”
Furthermore, Lawn et al. (1997) report on the “convergent evolution of apolipoprotein(a) in primates and hedgehog.” Apolipoprotein(a), or apo(a), a low-density lipoprotein (LDL) that is a risk factor for human atherosclerosis, is found in Old World primates (Catarrhini), and has also been identified in European hedgehogs. Remarkably, the protein functions identically in both organisms. The paper reports,
By apparent remodeling of a plasminogen-like gene, hedgehog and human ancestors independently evolved an apo(a) protein with multiple kringle domains that covalently links to apoB-containing lipoproteins, binds fibrin, lacks proteolytic activity, and competitively inhibits plasminogen activation.
An additional example is the “sequence convergence in the peptide-binding region of primate and rodent MHC Class Ib molecules” (Yeager et al., 1997) and the “independent origin of Prosimian, Platyrrhine, and Catarrhine Mhc-DRB genes” (Kriener et al., 2000). Moreover, Jost et al. (2008) report on “4 taxonomically diverse species of pufferfishes (Tetraodontidae)” which “each evolved resistance to the guanidinium toxins tetrodotoxin (TTX) and saxitoxin (STX) via parallel amino acid replacements across all 8 sodium channels present in teleost fish genomes.”
Even highly sophisticated molecular mechanisms have often evolved multiple times independently. One especially remarkable case of this is the convergent evolution of very similar DNA biosynthesis mechanisms in the archaea and the eubacteria (Leipe et al., 1999). Complex camera eyes have also arisen independently in multiple ineages, having evolved in chordates and molluscs, as well as alciopid annelid worms (Wald and Raypart, 1977) and two different groups of spiders (Laughlin, 1980; Williams and McIntyre, 1980).
Finally, one of the most astonishing cases of convergent evolution is discussed by Lukes et al. (2009). This paper concerns the evolution of two major distantly related phyla of flagellate protozoa, namely, the dinoflagellates and euglenozoa. These two groups are taxonomically so far apart that they are even members of different kingdoms (chromalveolates and excavates respectively). What is remarkable about these groups is that their evolutionary trajectories — indeed the evolution of “fundamental structures and processes” — have substantially deviated, in numerous respects, from other eukaryotes.
But here’s the point: Despite being very distantly related, these two lineages have deviated from the eukaryotic norm, in many cases, in much the same fashion. Indeed, there is a treasure trove of independently acquired features and characteristics. The paper (whose contents are helpfully summarized in this lecture) discusses numerous instances of convergent evolution pertinent to cellular organization, the nucleus, the plastid and the mitochondrion.
Convergent evolution is everywhere in biology. Many more examples could be given: The cases described above barely scratch the tip of the iceberg.
In his book Convergent Evolution: Limited Forms Most Beautiful, George McGhee reflects,
But what about the numerous cases in nature whether the morphological data and the nuclear data clash? In those cases, it is generally assumed that similarities in morphological traits have arisen by convergent evolution and are not synapomorphies, because the nuclear genome data can be trusted to be almost free from molecular convergences. What if this is not true?
In other words, extensive convergent evolution at the DNA level is considered to be so improbable under standard evolutionary assumptions that the nuclear data is preferred over morphological data in those many cases where conflict exists. In light of the numerous instances of demonstrable molecular convergence, however, the validity of this assumption — perhaps even some of the key tenets of the modern evolutionary paradigm — may need to be revisited. The widespread nature of homoplasy at both the molecular and morphological level substantially undercuts the frequent argument for common descent based on the nested hierarchical distribution of shared traits among organisms.
For further material on homoplasy and convergent evolution, in addition to George McGhee’s book mentioned above, I refer readers to Simon Conway Morris’s Life’s Solution: Inevitable Humans in a Lonely Universe (you can also find Conway Morris’s website on the topic here); as well as Fazale Rana’s contribution (chapter 21) to The Nature of Nature. No doubt the more genomes that are sequenced the more the pervasive nature of homoplasy will come to be appreciated.
Image credit: Wikipedia.

