 Intelligent Design
Intelligent Design
 Life Sciences
Life Sciences
Chemical Affinity, Motor Proteins: Good Questions from a Reader
A reader, Steve, posed a couple of good questions:
Dr. Gauger, thank you for taking my questions. I will try to be as short and to the point as possible.
First, regarding the aminoacylation of transfer RNA, is there any chemical bonding affinity inherent in the transfer RNA itself or does attachment of the amino acid to the 3′-end depend entirely upon energy provided by ATP? My thought is that if transfer RNA has no inherent chemical bonding affinity to any amino acid, in other words it is essentially dead in regard to chemical bonding, this constitutes a complete physical separation between the anticodon and the amino acid it represents. This makes the origin of the system via natural material forces extremely unlikely, especially when you add the fact that it depends on an irreducibly complex mechanism for its success.
Second, how do organelles, which are nonliving chemicals, appear to move with purpose? This question has two parts.
- How do organelles move from point A to point B? Cells will use cilia or flagella but I don’t see any mechanism causing a chaperone or a helicase to move.
- How do organelles move with purpose? My understanding is that organelles are non-living chemicals, yet they move in the cell as if they know what they are doing and where they are going. How is it that these non-living chemicals appear to have purpose and will move accordingly?
Thank you for taking the time to answer my questions. I am a Christian and I’m very interested in intelligent design theory. Your thoughts are deeply appreciated as they will help me to engage my friends and colleagues on this subject.
Steve seems to know something more than most about biology.
Questions and Answers
The match between the tRNA and its appropriate amino acid is made by specific enzymes called aminoacyl-tRNA synthetases. It is not done based on chemical affinity as some have tried to claim. Here is a wonderful page describing these enzymes and how they work:
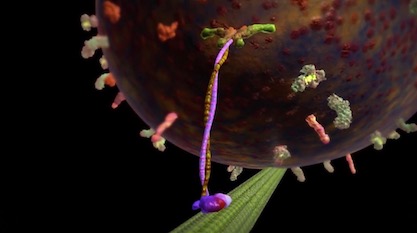
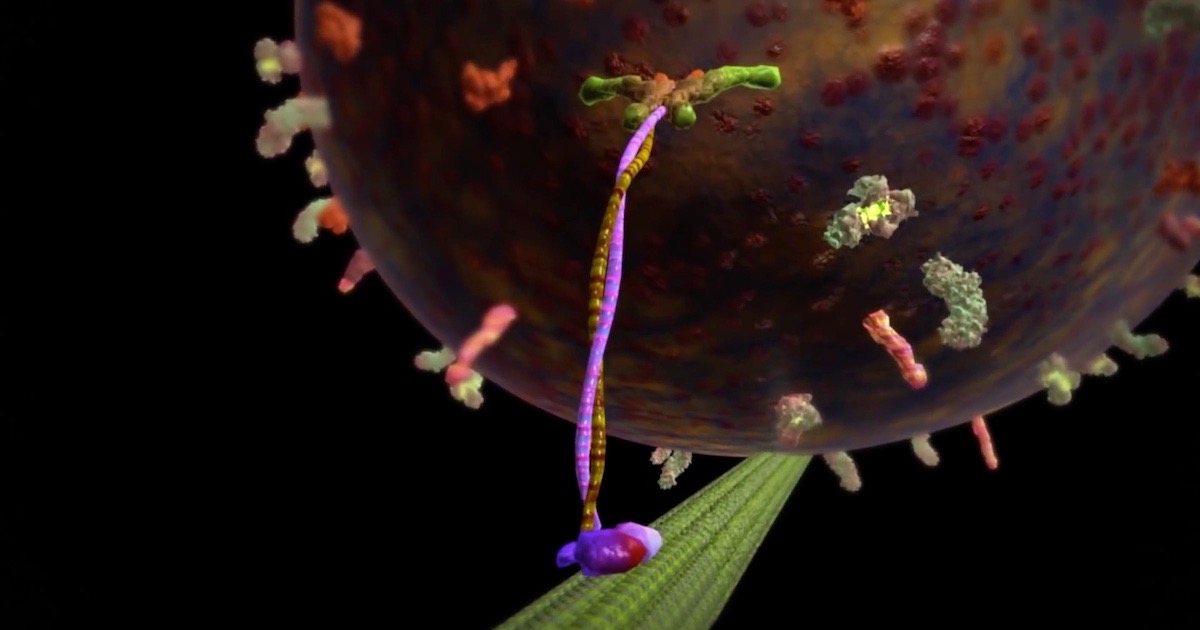
About the movement of organelles: There are molecular motor proteins that haul them around! For example, movement outward from the nucleus (called plus directed) is accomplished by a motor protein called kinesin, which runs on microtubes as a track. How they get directed to the right place is an active area of study and is what my postdoctoral work was on. Motion inward (minus directed) is accomplished by a motor protein called dynein. Dynein is involved in many critical processes in the cell. Here are a few videos and articles:
- “The Cell as a City: Examining an Apt Metaphor”
- “The Workhorse of the Cell: Kinesin”
- “Kinesin: Fast, Efficient, Essential, and Mysterious”
- “The Cell’s Postal System”
And there’s this about the “perfection” of molecular machines like kinesin:
And another wonderful video about the cell, with a really neat section about molecular motors at the very end.
I hope that helps.
Image: Kinesin, from The Workhorse of the Cell: Kinesin, via Discovery Institute.
