 Evolution
Evolution
 Intelligent Design
Intelligent Design
A Biochemical Icon of Intelligent Design, ATP Synthase Does More than Spin
The biochemical icon of rotary machines, ATP synthase, remains in the news with new discoveries. Now that cryo-electron microscopy is widely used, biophysicists are looking at the function of individual subunits of the engine and figuring out what they do. The fact that molecular machines exhibit more functional elegance the closer one looks indicates that intelligent design is the best explanation.
Automatic Brake
Animals and plants both contain these vital rotary engines that supply their energy needs in ATP. In animals, they are found in mitochondria. In plants and other photosynthetic organisms, they are found in chloroplasts. Photosynthesis, being dependent on light, has a problem: when it’s dark, their rotary engines might start running in reverse, risking “a wasteful ATP hydrolysis reaction.”
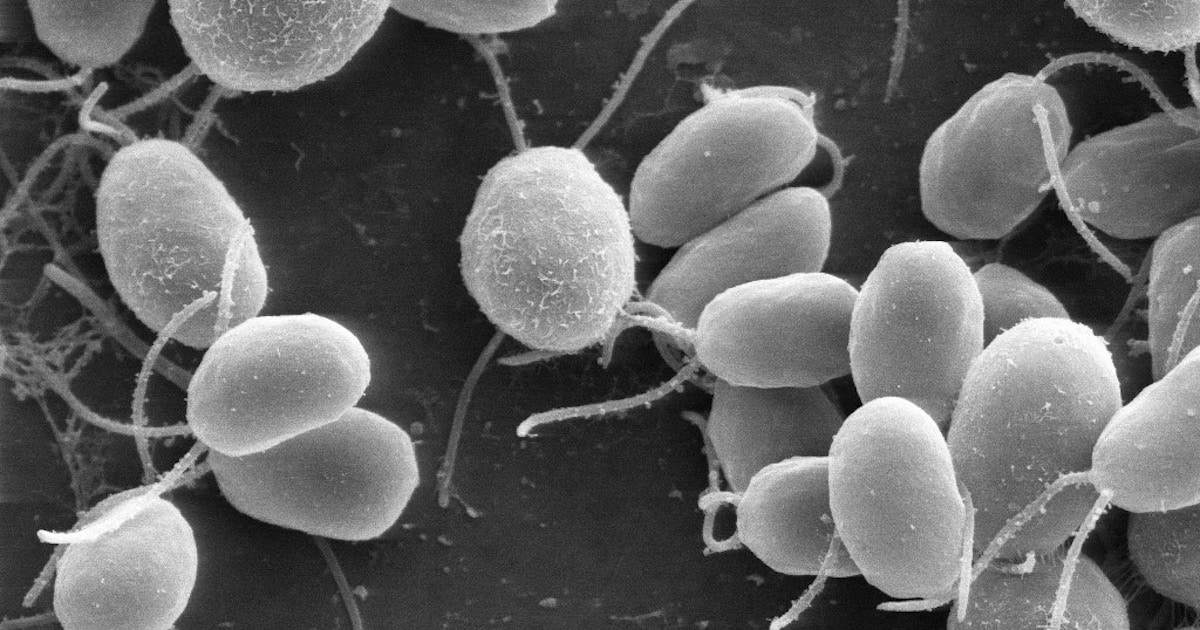
Working with the green alga Chlamydomonas reinhardtii (pictured above) as a “model organism,” a team of eight Japanese scientists took a closer look at the γ subunit in chloroplast ATP synthase (this is the “camshaft” part of the engine that drives the synthesis of ATP in the F1 domain). They found that two specific domains in the γ subunit act as an automatic brake in the dark. They wrote in PNAS in January,
Among the FoF1-ATP synthase complexes of all organisms, chloroplast FoF1 (CFoCF1) is a unique enzyme with a redox regulation mechanism; however, the underlying mechanism of redox regulation of the adenosine triphosphate (ATP) synthesis reaction in CFoCF1 has not been fully elucidated. By taking advantage of the powerful genetics of Chlamydomonas reinhardtii as a model organism for photosynthesis, we conducted a comprehensive biochemical analysis of the CFoCF1 molecule. Here we identify structural determinants for the kinetics of the intracellular redox response and demonstrate that the redox regulation of ATP synthesis is accomplished by the cooperative interaction of two γ subunit domains of CFoCF1 that are unique to photosynthetic organisms. [Emphasis added.]
Figure 6 in the article (reproduced by Phys.org) shows how the two domains act like a stopper:
The tight conformation weakens the interaction between the redox loop and the β-hairpin. Consequently, the β-hairpin remains stuck in the cavity between the α and β subunit, like a stopper, and inhibits the rotation of the central stalk (γεc-ring). In a reduced form, the redox loop recovers flexibility to interact with the β-hairpin. The redox loop interacts to pull out the β-hairpin from the cavity and thus accelerates the central stalk, like a cooperative regulator.
Watching the Snap
Another Japanese team, this one from the University of Tokyo, looked closer at the “catalytic dwell” in the F1 domain of the ATP synthase engine where ADP is converted to ATP with the addition of a phosphate. The F1 domain has 3 pairs of α, β subunits arranged like petals of a flower, each pair in a different phase of activity: insertion of ingredients, catalysis, and ejection of ATP. The γ subunit, like a camshaft, activates each α, β pair in turn as it rotates a full 360°. Dividing by three, each α, β pair feels the force of the camshaft during the catalysis stage (ADP + P yielding ATP) within 120° at each full turn of the crankshaft. At the other times, the pair is either receiving the ingredients or ejecting the finished ATP. When running in reverse, the F1 motor becomes an ATP cleaver, hydrolyzing ATP to yield ADP and P, ejecting protons in the process. In hydrolysis, then, ATP becomes the fuel to make the motor run in reverse.
During normal operation, the motor catalyzes ATP. Biophysicists have long suspected that the camshaft (the γ subunit) exerts pressure on the incoming ADP and P to snap them together. If 0° represents the moment ATP is catalyzed, previous studies have found a short pause stage at 80° and a longer dwell at the subsequent 40° of rotation, representing the “chemomechanical coupling scheme,” as their paper calls it.
In the reverse reaction, though, the angle for cleavage of ATP was unclarified. The team made a hybrid ATP synthase that ran extremely slowly so that they could observe “the world’s smallest rotary biological molecule motor” running in reverse. Their hybrid allowed them to measure the angle at which ATP cleavage occurs.
As a result, the new hybrid F1 showed two pausing angles that are separated by 200°. They are attributable to two slowed reaction steps in the mutated β, thus providing the direct evidence that ATP cleavage occurs at 200° rather than 80° subsequent to ATP binding at 0°. This scenario resolves the long-standing unclarified issue in the chemomechanical coupling scheme and gives insights into the mechanism of driving unidirectional rotation.
See the “Graphical Abstract” in the paper that illustrates this “extremely long dwell” they measured. The finding reveals that in both directions, ATP synthase is finely tuned for its work. The reverse direction (hydrolysis) is not just like a motor leaking fuel. Its parts act with precision to cleave ATP in a specific way.
Insights into a Related Rotary Motor
There’s a cousin to ATP synthase. It’s a proton pump called V-ATPase (vacuolar-type adenosine triphosphatase), and its catalytic parts are labeled V1Vo instead of F1Fo. Like its counterpart, V-ATPase runs with a rotary action but spends ATP to pump protons into organelles. Its job is to acidify vacuoles and other organelles or intracellular compartments that need a lower pH to work. The Vo part pumps the protons (H+ ions) into the vacuole, increasing its acidity. A little thought shows that such a motor could be dangerous. Would you want an acid-generating motor running loose?
Scientists from a Toronto hospital, publishing in PNAS, were curious to figure out how these acid pumps get built without causing harm to the cell. The Vo complex is assembled in the endoplasmic reticulum (ER) and then transported to the Golgi apparatus to be combined with V1. What quality control mechanism keeps the domains inactivated until they are fully assembled and ready for action?
Using cryo-electron microscopy, the team imaged three proteins (Vma12p, Vma22p, and Vma21p) that must work together to achieve the quality control for safely handling the acid pumps during assembly. “The resulting structures,” they found, “show how a sequence of coordinated interactions and conformational changes ensures that only properly assembled Vo leaves the ER and proton pumping into the neutral ER is avoided.” Spilling acid into the ER could be bad! “Unsurprisingly,” they remark, “owing to their importance for Vo assembly, mutations in the human homologues of Vma12p, Vma22p, and Vma21p have been linked to disease.” So how do these three essential proteins perform quality control? Do you really want to know?
The structures described above suggest the sequence of events that occur during Vo assembly in the ER membrane and subsequent binding of V1 in the Golgi…. The c ring assembles around Voa1p, with binding of subunit d to the c8c′c″Voa1p ring masking the ER-retrieval motif of Voa1p. The Vo∆aef:Vma12-22p structure indicates that the Vma12-22p complex binds this fully assembled ring prior to interaction with subunits a, e, and f (Fig. 4A). Vma12-22p helps recruit and secure the interaction of subunit a with the c ring through interaction of Vma12p with subunits a and d….
Their Final Sentence
OK, OK. Suffice it to say that a complicated set of interactions takes place to ensure V-ATPase assembly is safe! Biochemists may wish to labor through the details. Fortunately, the authors provided diagrams and animations to illustrate the dynamics of all these working parts. Note their final sentence: “Importantly, the structures illustrate how Vma21p and Vma12-22p play central roles in both V-ATPase assembly and quality control.”
Quality control: it’s an engineering concept that permeates all three of these studies. Without quality control, these nanoscopic rotary engines, on which all life depends, would never last — indeed, would never emerge in the first place. Quality control belongs in the working vocabulary of intelligent design and engineering. It is not found in Darwin’s dictionary.
