 Life Sciences
Life Sciences
Back to Basics: Reader Asks How We Know DNA Exists, What It Does
Here’s a thoughtful inquiry from a follower of Discovery Institute’s work, who recently asked us a basic but obviously important question: How do we know that DNA exists, and how do we know that it functions to carry genetic information? He also asks how we know what molecular machines like the nuclear pore complex look like.
Today, molecular biology is so advanced that it’s not every day that we think about these questions. As scientists push forward the frontiers of biological understanding, we take fundamental biological knowledge for granted. Nonetheless, it’s important to pause every so often and review how we know what we know.
Thankfully, with this in mind, I don’t need to give detailed answers because there are some great resources out there discussing these things. One excellent website that explains how we discovered DNA and genetics is DNA from the Beginning. This site, created by the famous Cold Spring Harbor Laboratory, lists dozens of steps through which “modern genetics” came to be. You’ll see how early thinkers and later scientists uncovered genetics and DNA, starting with rudimentary observations about how children resemble their parents, and later discovering that genes are carried by DNA, and ending with trying to understand non-coding DNA and epigenetic factors.
One of the most important steps occurred when scientists revealed that genes are made of DNA. Stephen Meyer tells the story in Chapter 4 of Signature in the Cell. Researchers injected DNA from dead bacteria into living ones, which were then injected into mice. The “dead DNA” helped the living bacteria produce a protein that allowed them to evade the mouse immune system, ultimately proving fatal to the mice involved. Chapter 5 in Signature in the Cell explains how biologists deduced the genetic code (including the processes of transcription and translation). These discoveries started off with an ultimately wrong — but not completely wrong — theoretical model by George Gamow, which it was then refined and improved by Francis Crick and many others.
X-ray crystallography also played a major role in illuminating the structure of DNA and many other important biological molecules, such as proteins. (The technique has helped us understand many inorganic compounds as well.) Today, X-ray crystallography isn’t as crucial to molecular biological research as it was in the past, but it was foundational to early studies of biological molecules. Basically, it works by crystallizing some compound and then bombarding it with X-rays. The angles at which the X-rays are diffracted and reflected off of electrons help us mathematically determine the structure and composition of the compound being analyzed. At Cold Spring Harbor Lab’s DNA Learning Center, you can see some great pictures showing how X-ray crystallography was used to determine the structure (and thus the function) of DNA.
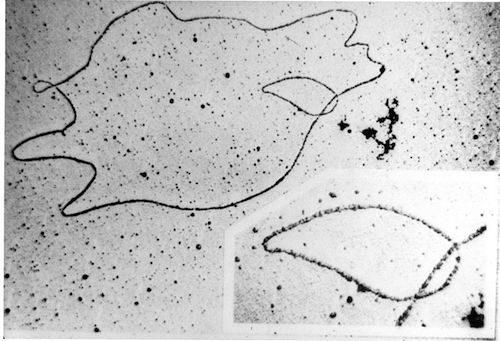
More recently, electron microscopy has revealed incredible images of a variety of molecular machines and cellular components whose fine structures are too small to see using light. That’s because the physical features of these machines are too small for even the tiny the wavelength of light to be able to distinguish. The wavelength of an electron is much smaller than that of a photon of light, so electron microscopy allows us to image much smaller features. We can even use this technique to directly take “pictures” of DNA, providing visual confirmation that DNA exists and showing what it looks like. For a nice article on this showing a photograph of the intense coiled packing of DNA, see “Scientists snap a picture of DNA’s double helix for the very first time” over at io9.
Electron microscopy has been applied to study the structure and function of many molecular machines, including the nuclear pore complex and eukaryotic flagella. Again, this technique provides direct visual evidence that these structures exist, shows what they look like, and can help us understand how they function.
Image: DNA under electron microscope, by SeanMack at en.wikipedia [Public domain], via Wikimedia Commons.
