 Intelligent Design
Intelligent Design
 Life Sciences
Life Sciences
Irreducible Complexity in Molecular Machine Assembly

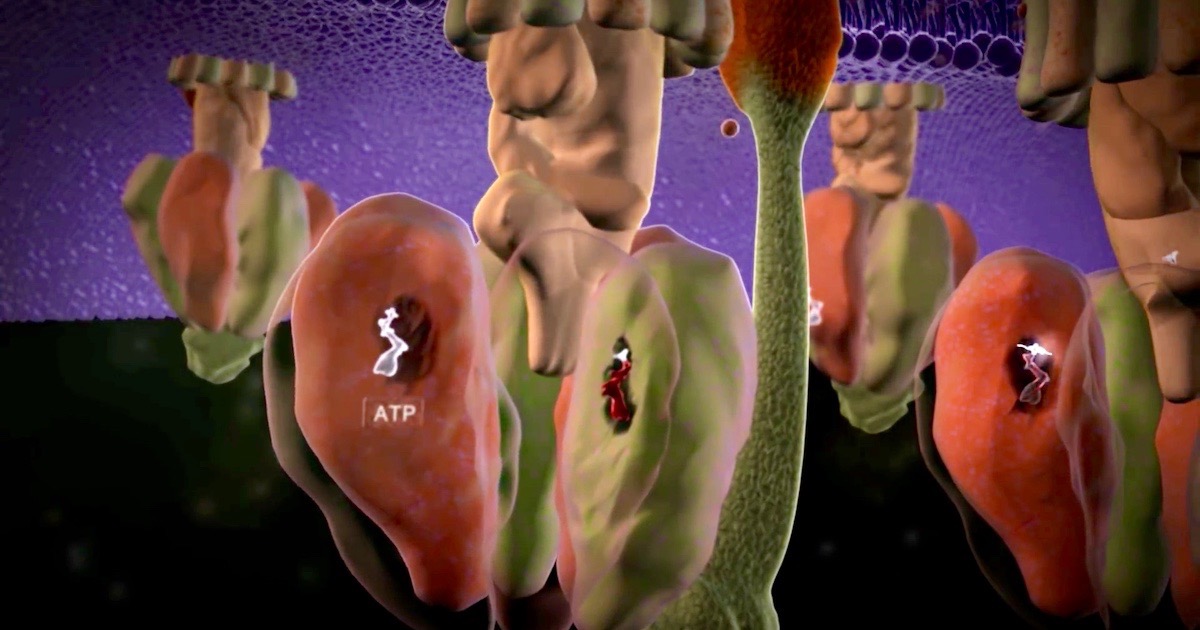
We know that many molecular machines are irreducibly complex (IC) in their operation. Even more IC is the process of assembling them in the cell. A good example of this is the process of building our good old standby machine, ATP synthase (review our animation to recognize the F0 rotating part and the F1 synthesis part).
A new “tour de force” paper by He et al. in the Proceedings of the National Academy of Sciences (PNAS), co-authored by Nobel laureate John E. Walker (who at age 77 is still researching these tiny rotary engines), describes new insights into how these multi-part machines are assembled. In a companion Commentary on PNAS, three scientists (Song, Pfanner and Becker) put it bluntly: “The assembly of the mitochondrial ATP synthase is a complicated process that involves the coordinated association of mitochondrially and nuclear encoded subunits.” Here’s a taste of what they mean (don’t worry; this won’t be on the test):
Based on their findings [He et al.], they propose an elegant model of how the membrane domain of human ATP synthase is built (Fig. 1, Upper). In one branch, an F1–c-ring intermediate associates with the peripheral stalk and the supernumerary subunits e and g. In the other branch, the F1 domain first assembles with the peripheral stalk and supernumerary subunits e, g, and f. Both pathways merge in a key assembly intermediate that contains the F1 domain, the c-ring, the peripheral stalk, and the supernumerary subunits e, g, and f. In all these vestigial [i.e., incomplete] ATP synthase complexes, the inhibitory protein IF1 is enriched to prevent ATP hydrolysis by the uncoupled ATP synthase. The presence of the supernumerary subunits e, g, and f is crucial for the subsequent integration of the mitochondrially encoded subunits ATP6 and ATP8 that are stabilized by addition of 6.8PL. Thus, the proton-conducting channel between ATP6 and the c-ring is formed. At this stage, ATP synthesis is coupled to the proton-motive force and the inhibitory protein IF1 is released. Finally, DAPIT is added to the assembly line to promote dimerization and oligomerization of the ATP synthase. [Emphasis added.]
Whether or not you can follow the jargon is not as important as what they witnessed: an “elegant” process that requires precise timing and coordination. Different machine parts must arrive on schedule, and assemble into intermediate (vestigial) forms that are nonfunctional alone. An inhibitor protein makes sure the machine doesn’t switch on ahead of schedule. The proton-conducting channel has to form just right so that it doesn’t “leak” protons. Only when all the parts are ready does the machine begin to rotate, but even then, the work isn’t complete. Another player is “added to the assembly line” to position the machines on the folds of the mitochondrial membrane (called cristae) at precise angles and spacings for optimum productivity.
The parts must arrive at the construction site on time. Some of them come from the nucleus, which must seem like many miles away at the scale of the machine. Some are built locally by genes within the mitochondrial genome. Interestingly, there are differences between yeast and humans regarding which genes are encoded where, and in what order they are assembled. But the proof of the pudding is in the respiration after eating: both versions of the machine work efficiently for their respective organisms.
The intermediate structure, somewhat like a scaffold on which the machine will be built, is also irreducibly complex:
We have shown that the assembly of human ATP synthase in the inner organellar membrane involves the formation of a monomeric intermediate made from 25 nuclear-encoded proteins into which the two mitochondrially encoded subunits are inserted and then sealed by association of another nuclear-encoded protein, thereby dimerizing the complex. Association of a final nuclear protein oligomerizes the dimers back-to-face along the cristae edges.
Notice that parts from the different genomes have to work tightly together. It’s like a manufacturing plant receiving parts locally and from India that have to meet agreed-on specifications to match. There are also rules for import, just like for parts arriving from a far country. The nuclear-encoded parts have to pass through two distinct checkpoints (the inner and outer membranes of the mitochondrion), which each have their robotic security personnel to validate them and facilitate their transport to the inside.
Previous work has shown how the completed “factory” of machines is organized within the mitochondrion. A specific nuclear protein seals them in two’s (dimers) at an angle, such that the rotating F0 proton pumps can maximize the intake of proton fuel, while the F1 parts, where ATP synthesis occurs, are farther apart to not crowd the output molecules. A “final nuclear protein” joins the dimers together (oligomerizes them) along the membrane edges. The longitudinal spacing is also tightly controlled, so that they don’t crowd each other. Every point of the assembly is programmatically directed. When everything is completed, rows of ATP synthase motors are arranged like turbines in a hydroelectric plant, feeding off a flow of protons produced by upstream machines in the respiration transport chain.
Ribosome Assembly
Viewers of cellular animations like those in Unlocking the Mystery of Life could never forget the assembly-line process inside the ribosome, where precisely-sequenced messenger RNAs are matched with transfer RNAs carrying amino acids to form proteins. The entrance tunnels for the ingredients and the exit tunnels for the polypeptides, and everything in between, must be positioned exactly for correct operation. The ribosome is certainly one of the most stunning examples of information translation in all of nature. But how is the ribosome itself built?
Nature has provided an early version of an unedited manuscript by Sanghai et al. about ribosome assembly. Although it has been accepted for publication, it will be subject to editorial revisions. The subject matter, though, appears to show another stunning case of irreducible complexity in the construction of this important molecular machine. Here’s the Abstract:
Early co-transcriptional events of eukaryotic ribosome assembly result in the formation of precursors of the small (40S) and large (60S) ribosomal subunits. A multitude of transient assembly factors regulate and chaperone the systematic folding of pre-ribosomal RNA subdomains. However, due to limited structural information, the role of these factors during early nucleolar 60S assembly is not fully understood. Here we have determined cryo-EM reconstructions of the nucleolar pre-60S ribosomal subunit in different conformational states at resolutions up to 3.4 Å. These reconstructions reveal how steric hindrance and molecular mimicry are used to prevent both premature folding states and binding of later factors. This is accomplished by the concerted activity of 21 ribosome assembly factors that stabilize and remodel pre-ribosomal RNA and ribosomal proteins. Among these factors, three Brix-domain proteins and their binding partners form a ring-like structure at rRNA domain boundaries to support the architecture of the maturing particle. Mutually exclusive conformations of these pre-60S particles suggest that the formation of the polypeptide exit tunnel is achieved through different folding pathways during subsequent stages of ribosome assembly. These structures rationalize previous genetic and biochemical data and highlight the mechanisms driving eukaryotic ribosome assembly in a unidirectional manner.
The requirements of IC are met in this description: “a multitude of transient assembly factors” regulate and systematically fold the proteins that will be used to construct the machine. The authors mention “21 ribosome assembly factors that stabilize and remodel” the RNA and proteins before the machine is even operational. Inside the growing ribosome, a scaffold holds factors for the exit tunnel in place. Everything is choreographed in time and space with “mechanisms driving… assembly in a unidirectional manner.”
Here we see numerous parts working together on a timeline. The parts alone do not work individually. You can have all the proteins delivered to the construction site, and nothing will happen without the programmed mechanisms to put them together in order. Some parts hold others in place, others guide the folding of protein parts, and some even prevent premature assembly. All the pathways for assembly of the subdomains are regulated by a master program, so that each group of steps follows a “unidirectional” plan toward the finished product. It’s a marvelous IC assembly process that produces an IC machine. If five parts of a mousetrap are sufficient to indicate IC, how about dozens of parts, all following a programmatic sequence of assembly?
In Unlocking, concerning the assembly of the bacterial flagellum, Paul Nelson described how the hierarchical IC of machine assembly in the cell challenges Darwinian theory. “In order to construct that flagellar mechanism, or tens of thousands of other such mechanisms in the cell, you require other machines to regulate the assembly of these structures. And those machines themselves require other machines for their assembly.” Jonathan Wells nailed the point by saying, “If even one of these pieces is missing, or put in the wrong place, your motor isn’t going to work. So this apparatus to assemble the flagellar motor is itself irreducibly complex. In fact, what we have here is irreducible complexity all the way down.”
Image source: ATP Synthase: The Power Plant of the Cell, via Discovery Institute.
