 Evolution
Evolution
 Intelligent Design
Intelligent Design
Can New Proteins Evolve?
Editor’s note: This excerpt is from a chapter by Douglas Axe in the newly released book The Comprehensive Guide to Science and Faith: Exploring the Ultimate Questions About Life and the Cosmos.
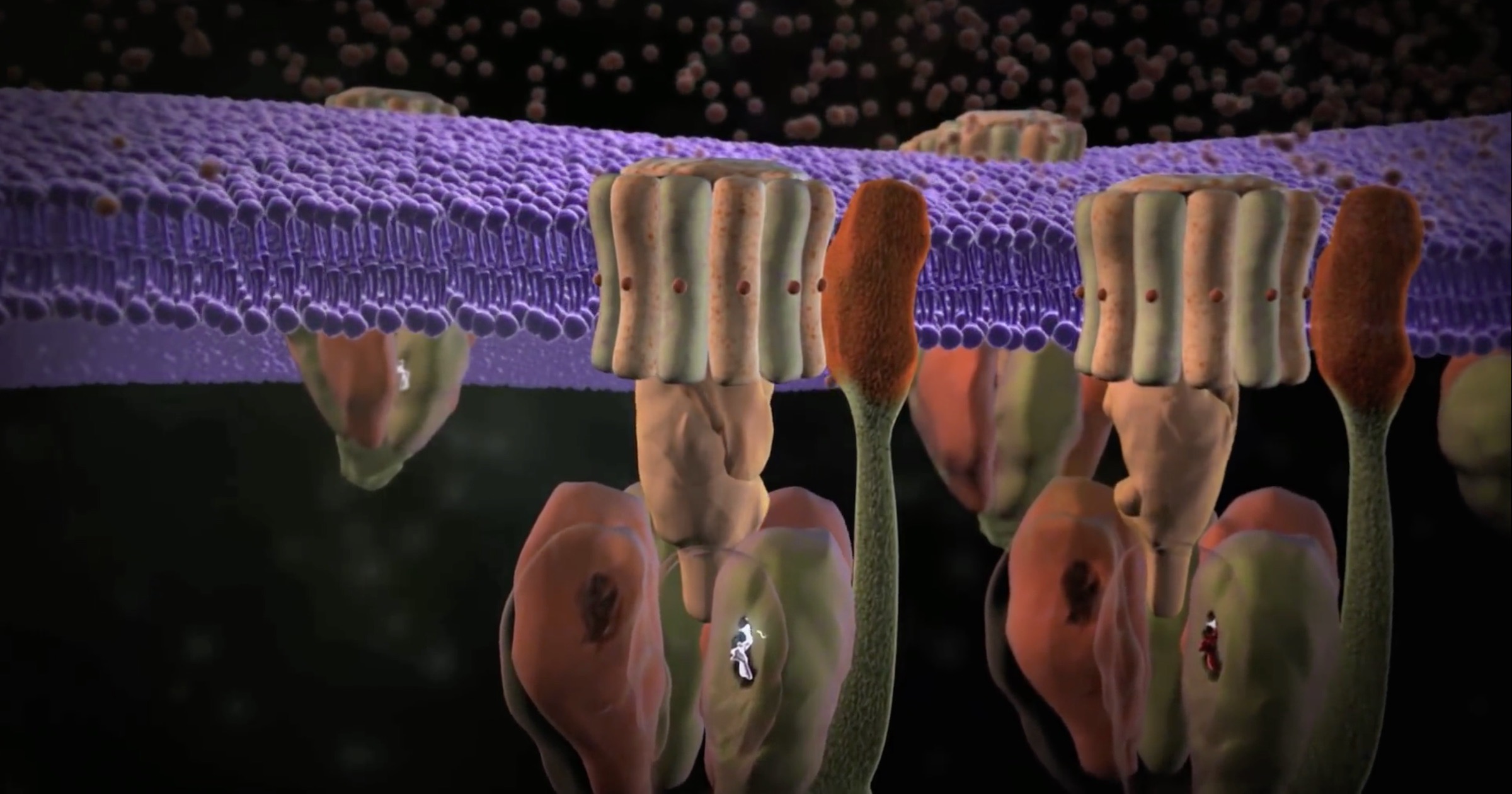
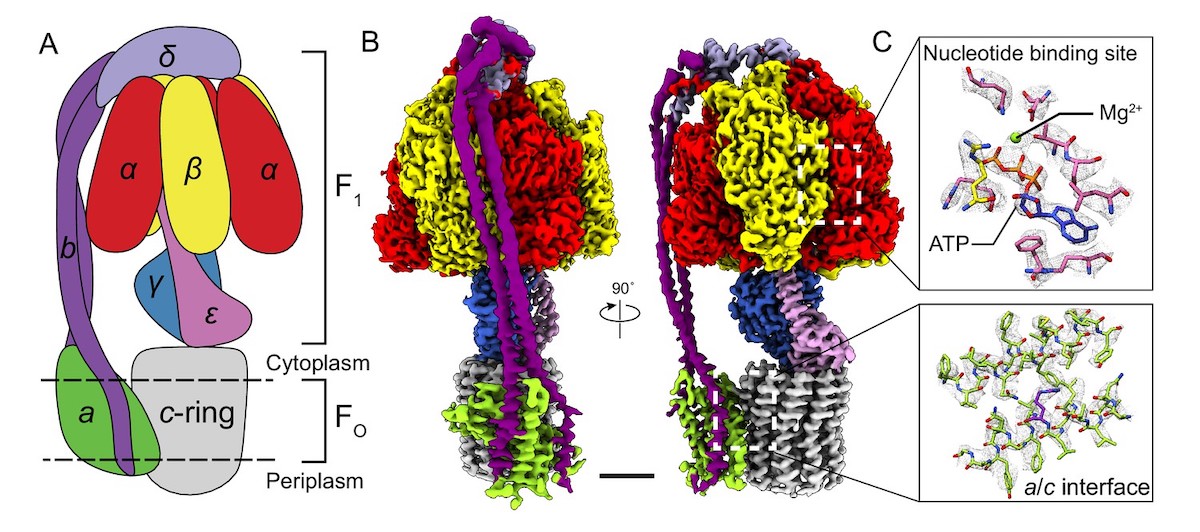
What enables a long chain of linked amino acids to perform highly specific molecular functions with machine-like precision? The answer is machine-like structure. Figure 2 shows one example (among thousands) of these remarkable structures. The multipart machine depicted is an ATP synthase — an assembly of 22 protein molecules that produces the energy molecule ATP. Biochemists refer to this as an enzyme because it accomplishes a chemical conversion (making ATP from ADP). But it’s no exaggeration to call it a molecular machine as well. Operating as a sophisticated nano-generator, the ATP synthase has a rotor (consisting of the parts labeled c-ring, y, and E) that spins at 8,000 rpm!
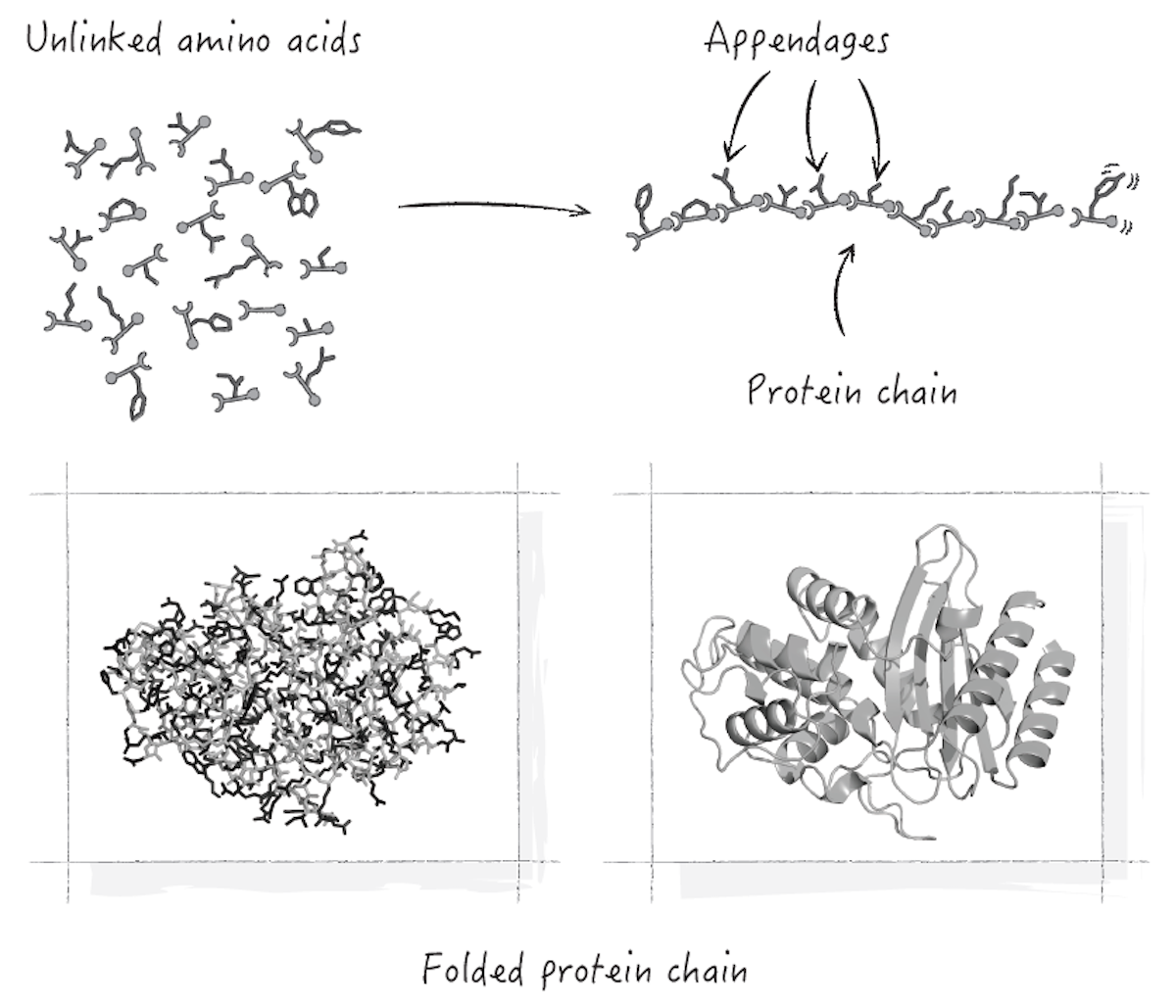
But how do long chains of linked amino acids form stable parts to make machines like this? The short answer is that it’s possible for the different amino acids to be arranged along the chain in such a way that the whole thing locks into a specific three-dimensional form. The process is called protein folding, with the term fold referring to the overall form. Figure 3 illustrates the basics.
I emphasized the word possible there for a reason. A random gene would specify a random sequence of amino acids, which would flop around without folding. Chains like that are rapidly broken back down into amino acids to keep them from interfering with cellular processes. Very special amino acid sequences are needed for protein chains to fold into stable structures.
Measuring the Remarkable
It’s possible to measure just how special these sequences must be. Because random genes are hopeless, the best way to do this is to start with a natural gene that specifies a selectable trait by producing a protein that imparts this trait. Laboratory methods exist for introducing random mutations into the chosen gene. This can be done in a controlled way by restricting how many changes occur or by confining the mutations to a small portion of the gene. By starting with a gene that specifies a working protein and by controlling in this way the extent to which the gene is mutated, experimenters can produce large numbers of mutant versions of the gene, some of which will almost certainly still work.
Building on earlier work of this kind, I applied the method to a natural gene that enables bacteria to inactivate penicillin-like antibiotics. The trait in this case is antibiotic resistance, which is very easy to select for in the lab. First, one simply puts the bacteria on petri dishes with a small amount of penicillin. Cells carrying a mutant version of the gene specifying an enzyme that’s still able to break penicillin will form visible colonies on the petri dishes, whereas cells with inactive genes will die. I chose four clusters of amino acids, ten each, for my experiments. In each experiment, I heavily mutated the gene locations for one cluster, resulting in many mutant genes, each of which specified a mutant version of the enzyme with a jumble of amino acids in those ten locations. Of roughly 100,000 mutant genes tested per cluster, some were found to work in three of the clusters. None in the fourth. Testing more mutants presumably would have turned up some that worked in that fourth cluster. In any case, I was able to estimate from these results a fraction of mutants that work for each cluster, though this fraction is really an upper-bound estimate for the fourth cluster.
From these experimentally based estimates, I calculated the fraction of mutants that would be expected to work if the entire gene had been mutated in a similar way. This fraction is closely related to another fraction that carries a great deal of significance for protein evolution. But before we talk about what this important fraction turned out to be, let’s take some time to understand evolutionary thinking well enough to put this fraction in its proper context…..
Back to Proteins
Returning now to proteins, what do they add to the picture? The answer is that they exemplify the commonsensical principles by which we rightly reject purpose-free accounts of life. We can fully grasp and affirm these principles without turning to the subject of proteins (or any other technical subject), but in proteins we find elegant confirmation.
The important fraction I referred to above is the likelihood that a random chain of amino acids would have the special physical properties needed to fold into a stable structure that’s suitable for a particular function. Putting that in evolutionary terms, assuming a particular new capability that can be achieved with a new protein fold would benefit an organism, and that a genetic mistake in that organism has produced a gene sequence that differs substantially from what existed before, this fraction is the probability that the new gene happens to encode a new protein that performs the desirable new function. Using the results of my experiments on the penicillin-resistance enzyme, I estimated this probability to be in the ballpark of:
1/100,000,000,000,000,000,000,000,000,000,000,000,000,000,000,000,000,000,000,000,000,000,000,000,000,000
