 Evolution
Evolution
 Life Sciences
Life Sciences
Asking the Right Questions about the Evolutionary Origin of New Biological Information
| Links to our 8-Part Series, “The NCSE, Judge Jones, and Citation Bluffs About the Origin of New Functional Genetic Information”: • Part 1: Judge Jones’s Misguided NCSE-Scripted Kitzmiller Ruling and the Origin of New Functional Genetic Information Read the Full Article: “The NCSE, Judge Jones, and Citation Bluffs About the Origin of New Functional Genetic Information” |
As we’ve seen, it’s easy to duplicate a gene, but the key missing ingredient in many neo-Darwinian explanations of the origin of new genetic information is how a gene duplicate then acquires some new optimized function. Evolutionists have not demonstrated, except in rare cases, that step-wise paths to new function for duplicate genes exist.
As we saw in an earlier post, Austin Hughes cautions against making “statistically based claim[s] of evidence for positive selection divorced from any biological mechanism.”26 In other words, natural selection is invoked to explain the evolution of genes where we do not even know the functional effect of the mutations being asserted. In this regard, Hughes observes that even in one of the more sophisticated studies, “there was no direct evidence that natural selection was actually involved in fixing adaptive changes.”27
Hughes also acknowledges a problem inherent in many appeals to natural selection, namely that required mutations may not give any selective advantage when they first arise. He thus writes regarding one study:
For example, a rhodopsin from the Japanese conger eel with λmax ≈ 480 nm achieved this sensitivity through the interaction of three different amino acid replacements (at sites 195, 195, and 292). There does not seem to be any way that natural selection could favor an amino acid replacement that would be of adaptive value only if two other replacements were to occur as well.28
In this case, there was no stepwise advantage gained with each successive mutation. Because no advantage could have been gained until all three mutations were present, Hughes finds it more “plausible” to believe that the first two mutations were “selectively neutral” and became fixed due to random, non-adaptive processes such as genetic drift. Once the third mutation arose it might have provided an advantage, but to paraphrase Scott Gilbert, at best this really only explains the survival of the fittest, not the arrival of the fittest.29
But Hughes’ explanation has deep deficiencies: it requires that two mutations become fixed before any selective advantage for the third mutation is gained. This implies that there must be three specific mutations to gain any selective advantage. A key question is thus, Are multiple specific mutational changes likely to appear in the same individual through unguided chance mutations given known mutation rates and population sizes? Even Hughes, despite his exhortations to fellow evolutionary biologists to employ more rigor in their studies, does not address this fundamental question.
A similar example is found when leading paleoanthropologist Bernard Wood critiqued a simplistic model of human cranial evolution on the grounds that too many mutations would be required to gain any functional advantage:
The mutation would have reduced the Darwinian fitness of those individuals. . . . It only would’ve become fixed if it coincided with mutations that reduced tooth size, jaw size and increased brain size. What are the chances of that? 30
Similarly, Jerry Coyne writes that “It is indeed true that natural selection cannot build any feature in which intermediate steps do not confer a net benefit on the organism.”31 This highlights a key deficiency in many neo-Darwinian accounts of the evolution of genes. Namely, they fail to demonstrate that the processes necessary to generate new functionally advantageous genetic information are plausible. As with Hughes’s or Wood’s examples above, multiple mutations might be necessary to gain any functional advantage. Any account invoking blind, unguided, random mutations to evolve a gene from Function A to Function B must address at least these three questions:
- Question 1: Is there a step-wise adaptive pathway to mutate from A to B, with a selective advantage gained at each small step of the pathway?
- Question 2: If not, are multiple specific mutations ever necessary to gain or improve function?
- Question 3: If so, are such multi-mutation events likely to occur given the available probabilistic resources?
Mathematician David Berlinski considers such questions when critiquing evolutionary accounts of eye evolution. Darwinian processes fail because multiple changes are required for a new function to appear:
If these changes come about simultaneously, it makes no sense to talk of a gradual ascent of Mount Improbable. If they do not come about simultaneously, it is not clear why they should come about at all.32
Again, the key question is therefore, how hard is it for new functional biological information to arise? Answering this question requires assessing the ability of random mutation and natural selection to generate new functional biological information. But when most evolutionary biologists play the Gene Evolution Game, they do not make such assessments and rarely consider these questions. Instead they typically invoke processes such as gene duplication, natural selection, and rearrangement, without demonstrating that random and unguided mutations are sufficient to produce the information needed. Any explanation that at base is little more complicated than “duplication, rearrangement, and natural selection” is not a demonstration that new functional genes can arise by unguided processes.
Thankfully, some scientists are willing to consider these key questions. They have performed research providing data that offers strong reasons to be skeptical of the ability of mutation and selection to form new functional genetic sequences.
A. Asking Questions 1 and 2:
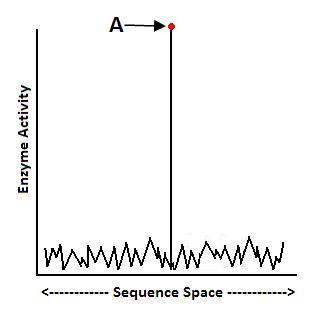
Molecular biologist Doug Axe has performed mutational sensitivity tests on enzymes and found that functional protein folds may be as rare as 1 in 1077.33 His research shows that the fitness landscape for many enzymes looks like this, making it very unlikely that neo-Darwinian processes will find the specific amino acid sequences that yield functional protein folds:
To put the matter in perspective, these results indicate that the odds of Darwinian processes generating a functional protein fold are less than the odds of someone closing his eyes and firing an arrow into the Milky Way galaxy, and hitting one pre-selected atom.34 To say the least, this exhausts the probabilistic resources available. Such data help us answer the first question: it’s not likely that there will be a functional stepwise mutational pathway leading from Function A to Function B.
Douglas Axe is by no means the only biologist to make this observation. A leading college-level biology textbook, Campbell’s Biology, observes that “Even a slight change in primary structure can affect a protein’s conformation and ability to function.”35 Likewise, David S. Goodsell, an evolutionist biologist, writes:
As you might imagine, only a small fraction of the possible combinations of amino acids will fold spontaneously into a stable structure. If you make a protein with a random sequence of amino acids, chances are that it will only form a gooey tangle when placed in water. Cells have perfected the sequences of amino acids over many years of evolutionary selection…36
What Goodsell does not mention is that if “perfected” amino acid sequences and functional protein folds are rare and slight changes can disrupt function, then selection will be highly unlikely to take proteins from one functional fold to the next without traversing some non-functional stage. So how do new functional protein folds evolve? This effectively answers question two, implying that many specific mutations would be necessary for evolving genes to pass through non-functional stages while evolving some new function. Question 3 assesses whether this is likely to happen.
B. Asking Question 3:
In 2004, Michael Behe and physicist David Snoke published a paper in the journal Protein Science reporting results of computer simulations and theoretical calculations. They showed that the Darwinian evolution of a simple functional bond between two proteins would be highly unlikely to occur in populations of multicellular organisms. The reason, simply put, is because too many amino acids would have to be fixed by non-adaptive mutations before gaining any functional binding interaction. They found:
The fact that very large population sizes–109 or greater–are required to build even a minimal [multi-residue] feature requiring two nucleotide alterations within 108 generations by the processes described in our model, and that enormous population sizes are required for more complex features or shorter times, seems to indicate that the mechanism of gene duplication and point mutation alone would be ineffective, at least for multicellular diploid species, because few multicellular species reach the required population sizes.37
According to this data, chance mutations are unlikely to produce even two required non-adaptive mutations in multicellular diploid species within any reasonable timescale. This answers the third question: getting multiple specific non-adaptive mutations in one individual is extremely difficult, and more than two required but non-adaptive mutations are likely beyond the reach of multi-cellular organisms. Studies like this show that the actual ability of random mutation and unguided selection to produce even modestly complex new genetic functions is insufficient.
In 2008, Behe and Snoke’s would-be critics tried to refute them in the journal Genetics, but found that to obtain only two specific mutations via Darwinian evolution “for humans with a much smaller effective population size, this type of change would take > 100 million years.” The critics admitted this was “very unlikely to occur on a reasonable timescale.” 38 In other words, there is too much complex and specified information in many proteins and enzymes to be generated in humans by Darwinian processes on a reasonable evolutionary timescale.
As noted in the comments on the Gene Evolution Game, when neo-Darwinists try to explain the evolution of genes, mere point mutations often are insufficient to account for the gene’s sequence. They must therefore appeal to genetic rearrangements such as insertions, deletions, or an alleged process called “domain shuffling” where segments of proteins become shuffled to new positions in the genome. In his book The Edge of Evolution, Michael Behe reviews research that engineered new protein function by swapping domains to change protein function, and found that the intelligently engineered changes required multiple modifications that, in nature, would require too many simultaneous mutational events to yield functional changes:
[Protein engineering research] does not mimic random mutation. It is the exact opposite of random mutation. … What do the lab results tell us about whether random-yet-productive shuffling of domains “occurs with significant frequency under conditions that are likely to occur in nature”? About whether that is biologically reasonable? Nothing at all. When a scientist intentionally arranges fragments of genes in order to maximize the chances of their interacting productively, he has left Darwin far, far behind. … [Experiments that engineered proteins by shuffling domains] didn’t just splice two genes together in a single step; they took several additional steps as well. … Remember the more steps that have to occur between beneficial states, the much less plausible are Darwinian explanations. … Domain shuffling would be an instance of the “natural genetic engineering” championed by James Shapiro where evolution by big random changes is hoped to do what evolution by small random changes can’t. But random is random. No matter if a monkey is rearranging single letters or whole chapters, incoherence plagues every step. … One step might luckily be helpful on occasion, maybe rarely a second step might build on it. But Darwinian processes in particular and unintelligent ones in general don’t build coherent systems. So it is biologically most reasonable to conclude that, like multiple brand new protein-protein binding sites, the arrangement of multiple genetic elements into sophisticated logic circuits similar to those of computers is also well beyond the edge of Darwinian evolution. 39
As Behe observes, “No matter if a monkey is rearranging single letters or whole chapters, incoherence plagues every step.” Thus, when multiple mutational events–whether point mutations, “domain shuffling,” or other types of rearrangements–are required to gain some functional advantage, it seems unlikely that blind neo-Darwinian processes can produce the new biological function.
Unfortunately, few if any advocates of the neo-Darwinian just-so stories investigate whether mutation and natural selection are sufficient to produce new functional genetic information. Instead they believe that finding similarities and differences between genes demonstrates that neo-Darwinian evolution has occurred, and they assume that “positive selection” is a sufficient explanation.
As Hughes cautions, they engage in “use of certain poorly conceived statistical methods to test for positive selection,” causing “the literature of evolutionary biology [to become] glutted with extravagant claims of positive selection” resulting in a “vast outpouring of pseudo-Darwinian hype [that] has been genuinely harmful to the credibility of evolutionary biology as a science.” 40 Or, as Michael Behe cautions, they confuse mere sequence similarity with evidence of neo-Darwinian evolution. Finally, Michael Lynch warns his colleagues that “Evolutionary biology is not a story-telling exercise, and the goal of population genetics is not to be inspiring, but to be explanatory.” 41
With these principles in mind, in the next installment we will assess about a dozen of the just-so stories concerning the origin of genes offered in studies cited by the NCSE.
References Cited:
[26.] Austin L. Hughes, “Looking for Darwin in all the wrong places: the misguided quest for positive selection at the nucleotide sequence level,” Heredity, Vol. 99:364–373 (2007).
[27.] Id.
[28.] Id.
[29.] “The modern synthesis is good at modeling the survival of the fittest, but not the arrival of the fittest.” Scott Gilbert, quoted in John Whitfield, “Biological Theory: Postmodern evolution?,” Nature, Vol. 455:281-284 (2008).
[30.] Bernard Wood, quoted in Joseph B. Verrengia, “Gene Mutation Said Linked to Evolution,” Associated Press, found in San Diego Union Tribune, March 24, 2004.
[31.] Jerry Coyne, “The Great Mutator,” The New Republic (June 14, 2007). Coyne asserts he knows of no example where this is the case.
[32.] David Berlinski, “Keeping an Eye on Evolution: Richard Dawkins, a relentless Darwinian spear carrier, trips over Mount Improbable. Review of Climbing Mount Improbable by Richard Dawkins (W. H. Norton & Company, Inc. 1996),” in The Globe & Mail (November 2, 1996) at http://www.discovery.org/a/132
[33.] Douglas D. Axe, “Estimating the Prevalence of Protein Sequences Adopting Functional Enzyme Folds,” Journal of Molecular Biology, Vol. 341: 1295-1315 (2004); Douglas D. Axe, “Extreme Functional Sensitivity to Conservative Amino Acid Changes on Enzyme Exteriors,” Journal of Molecular Biology, Vol. 301: 585-595 (2000).
[34.] See Stephen C. Meyer, Signature in the Cell: DNA and the Evidence for Intelligent Design, pg. 211 (Harper One, 2009).
[35.] Neil A. Campbell and Jane B. Reece, Biology, pg. 84 (7th ed, 2005).
[36.] David S. Goodsell, The Machinery of Life, pg. 17, 19 (2nd ed, Springer, 2009).
[37.] Michael J. Behe & David W. Snoke, “Simulating Evolution by Gene Duplication of Protein Features That Require Multiple Amino Acid Residues,” Protein Science, Vol 13:2651-2664 (2004).
[38.] Rick Durrett and Deena Schmidt, “Waiting for Two Mutations: With Applications to Regulatory Sequence Evolution and the Limits of Darwinian Evolution,” Genetics, Vol. 180: 1501–1509 (November 2008).
[39.] Michael Behe, The Edge of Evolution: The Search for the Limits of Darwinism, Appendix D, pgs. 272-275 (Free Press, 2007) (emphasis added).
[40.] Austin L. Hughes, “The origin of adaptive phenotypes,” Proceedings of the National Academy of Sciences USA, Vol. 105(36):13193–13194 (Sept. 9, 2008) (internal citations removed).
[41.] Michael Lynch, “The frailty of adaptive hypotheses for the origins of organismal complexity,” Proceedings of the National Academy of Sciences, Vol. 104:8597–8604 (May 15, 2007).
