 Evolution
Evolution
 Intelligent Design
Intelligent Design
From Winston Ewert, New Peer-Reviewed Paper on Dependency Graph Model

Winston Ewert has published a new paper in the journal BIO-Complexity expanding on his previous research related to the pattern of similarities between species. He further demonstrated how the distribution of similarities fits his dependency graph model better than the standard common ancestry model. The paper can be downloaded here. I described the superior explanatory power of Ewert’s model here. His original BIO-Complexity paper is here.
The Dawkins Test
One of the central predictions of evolutionary theory is that similarities between species should consistently point to the same evolutionary tree based on the premise of common ancestry. The branching points in the tree should correspond to the common ancestor of a group of organisms (e.g., mammals) that first acquired all the traits common to that group (e.g., glands to produce milk and three bones in the inner ear).
Richard Dawkins stated that a reliable test of the design hypothesis for life is whether similarities are highly inconsistent with a single tree. Data collected over the past several decades has demonstrated that the distribution of similarities is far more inconsistent with common ancestry than any evolutionist imagined, so the intelligent design framework has passed the Dawkins test. Casey Luskin cited numerous studies detailing the inconsistencies here.
This data undermines evolutionary theory at its core. Nearly all the arguments for the grand evolutionary narratives (e.g., transformation of fish into humans) rest on the assumption that similarities reliably point to common ancestry. In many groups, the high level of inconsistency of similarities with any evolutionary tree demonstrates that this foundational assumption is false.
Particularly problematic, the places where evolutionists most require reliable data is where the data is most inconsistent. For instance, a central icon of evolution is the “whale series” consisting of fossils that purportedly illustrate a land mammal incrementally transforming into a fully aquatic mammal. The animals in the series are not believed to have directly descend from or evolved into each other, but their similarities are interpreted as their having descended from animals that are part of an ancestor-descendent series encompassing the land-sea transition.
The challenge is that the rescaled consistency index (RCI) for aquatic mammals and their purported ancestors is 0.24. An RCI value of 1.0 corresponds to perfect consistency of all traits with one evolutionary tree, and a value of 0.0 corresponds to traits that are randomly distributed. An RCI value significantly lower than 0.5 indicates that similarities are so inconsistent with the assumption of common ancestry that arguments for evolution based on similarities, such as with the whale series, become highly suspect. As another key example, the RCI for primates and their purported relatives is 0.29, so claims related to the evolution of humans resting on similarities in fossils also carry little weight.
Dependency Graph Model
In contrast, the pattern of similarities fits design-based models, as I previously described here. To summarize, human engineers use design modules, such as engines, in different creations to meet common goals. Likewise, similarities between species are often not consistent with an evolutionary tree, but they appear implemented in different creatures for a common purpose. Ewert’s framework is the first to identify modules based on proteins.
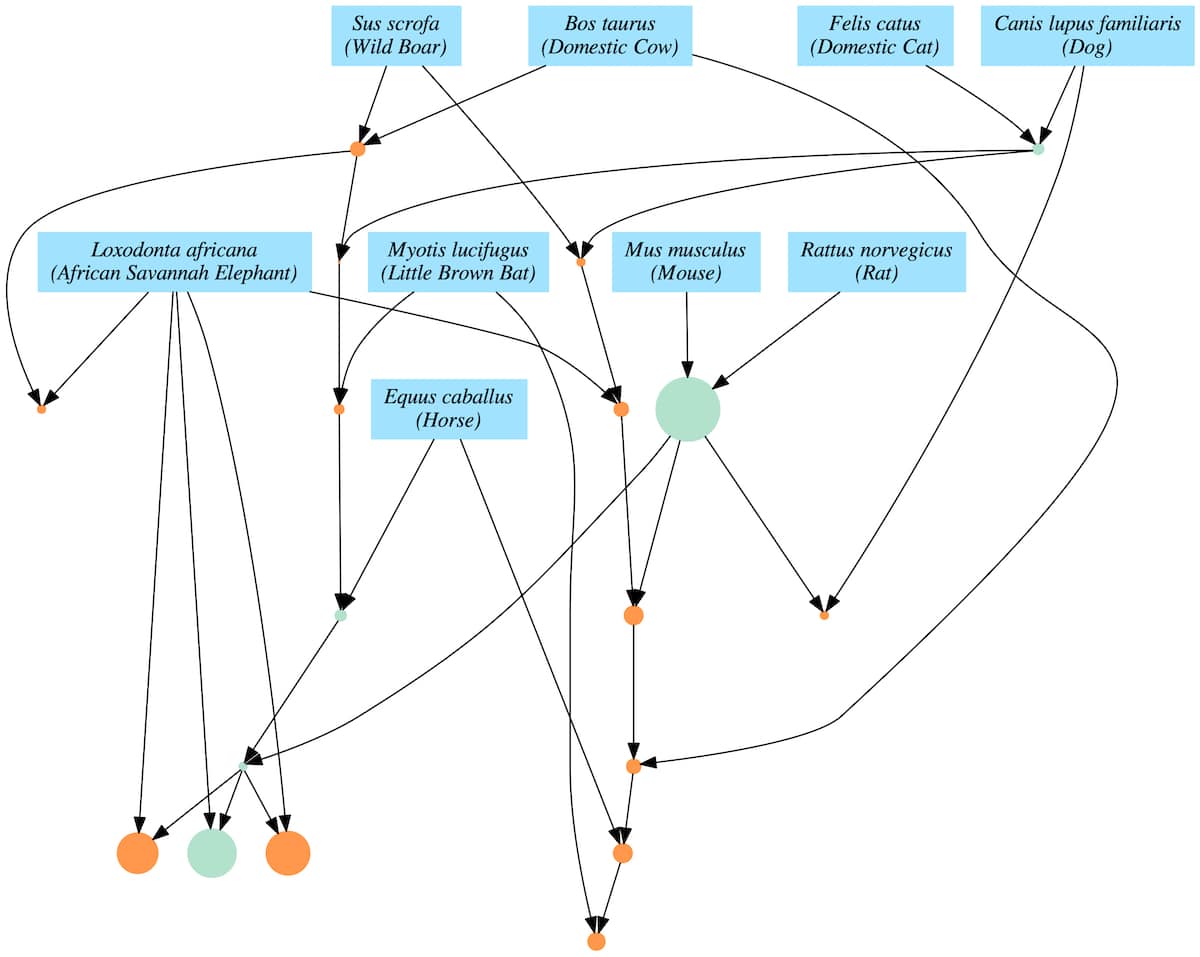
In Ewert’s first paper, he identified modules as sets of proteins from distinct families. Some modules are dependent on smaller modules, and the smaller modules are at times used by multiple larger modules (Figure 1). Ewert demonstrated quantitatively that the protein data far better fits the dependency graph model than common ancestry.
Application to Protein Sequences
In Ewert’s recently published article, he did not identify modules as sets of distinct proteins. Instead, he focused on variations in the protein prestin between a variety of mammal species including those capable of echolocation. The protein plays a critical role in mammal hearing by serving as a motor protein in the outer hair cells. The versions of the protein supporting echolocating bats, whales, and dolphins show the same amino acid alterations compared to the versions in mammals lacking echolocation. The pattern of alterations is not simple. Different pairs of mammals share similar amino acid alterations. Constructed evolutionary trees for mammals based on common ancestry lose this information while Ewert’s model highlights it.
To visualize the interrelationships, Ewert created a program called AminoGraph that identifies modules as specific amino acid alterations from an archetypal sequence that is a representative version of the protein. One module depends on another if it includes the latter’s alterations plus additional alterations. The program takes as input amino acid sequences for the same protein in different species, and it creates from this data the most consistent graph of module dependences.
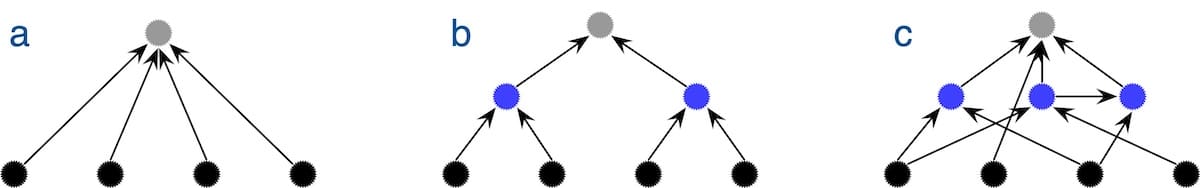
The program displays sets of sequences that are not related as each input sequence directly linking to the archetypal sequence (Figure 2a). It displays data that best fits a common ancestry model as a standard evolutionary tree (Figure 2b), and it displays data that corresponds to modules with complex interrelationships as a dependency graph (Figure 2c). Ewert tested the program on simulated data corresponding to unrelated sequences, sequences connected by common ancestry, and sequences connected through a specific dependency graph. In each case, the program properly identified the correct structure and accurately identified most of the modules and their interrelationships, thus confirming the reliability of the program.
Results for Prestin Protein
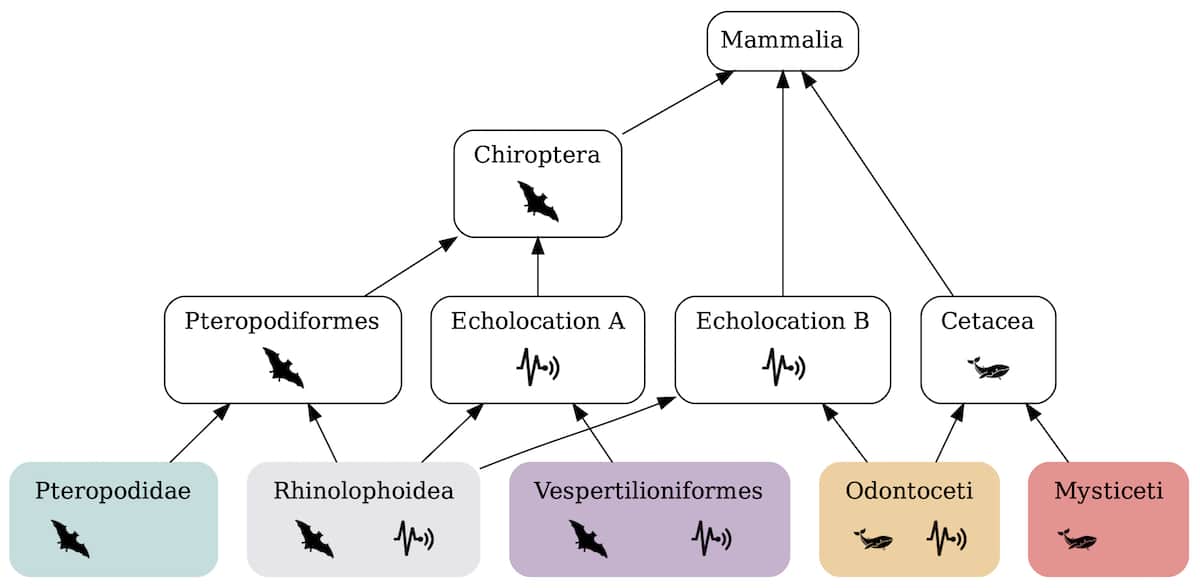
Ewert ran AminoGraph on the sequences of prestin proteins from species of echolocating bats, bats without echolocation, echolocating whales, whales without echolocation, and other mammals without echolocation. The generated dependency graph revealed a key insight into how the prestin proteins were modified to respond to the high sound frequencies generated in echolocation. The reengineering in each echolocating mammal employed some combination of two modules labeled Echolocation A and Echolocation B (Figure 3).
Future Research
This result naturally leads to research into the effects of amino acid modifications. The design framework predicts that alterations coordinate in each module to support echolocation in a way that best serves the needs of the species that employ it. Another expectation is that the prestin modules must coordinate with other modules for echolocation to function at a level that would benefit its possessor. This prediction appears consistent with the numerous similarities between echolocating mammals in both their genetics (here, here) and their overarching design logic (here, here).
In contrast, the standard evolutionary framework provides no insight into how echolocation is implemented in different mammals. The commonly accepted evolutionary tree requires echolocation to have evolved independently three times as illustrated in Ewert (2023), Figure 1. Yet the maximum possible time for the evolution of a fully aquatic mammal or a bat is insufficient for more than two coordinated mutations to appear. Any evolutionary scenario for echolocation would require far more than two coordinated mutations (here,here), so evolution fails to explain this trait’s origin even once, let alone multiple times.
Ewert’s model could be applied to several other proteins in echolocation and in other biological systems to gain similar insights. It represents a valuable tool in the developing theory of biological design, which should eventually supplant phylogenetic analyses.
